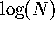 generations. This is much less than 2N generations which is the expected
time until fixation under selective neutrality with the Poisson progeny
distribution.
generations. This is much less than 2N generations which is the expected
time until fixation under selective neutrality with the Poisson progeny
distribution.
Heuristically, the coalescent process is the time reversal of the fixation process. This relation does not hold exactly because several successive generations may first attain fixation of one of their genes in the same generation, and several successive generations may share the same most recent common ancestor. But it is true that the average coalescent time (time since a common ancestor of the entire population) and average fixation time will be the same.
Indeed, the expected time since a common ancestor and expected time until fixation for the canonical selectively neutral random mating population with N diploid individuals and a Poisson progeny distribution with parameter 1 are both 4N generations (half that if the population has N haploid individuals). In general, we may use the diffusion approximation for fixation time to determine the expected coalescent time.
Progeny Distribution and Selection
Change in allele frequencies requires variation in progeny production. The
variance of the progeny distribution provides a bound on the rate at which
gene substitution can occur. If the progeny distribution is Poisson with
parameter 1, a single gene can become fixed in the population in 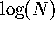 generations. This is much less than 2N generations which is the expected
time until fixation under selective neutrality with the Poisson progeny
distribution.
generations. This is much less than 2N generations which is the expected
time until fixation under selective neutrality with the Poisson progeny
distribution.
An alternative definition of the intensity of selection in a population is the autocorrelation between the number of progeny an individual has and the number of progeny its parent had. (The prefix auto refers to the fact that the correlation is within lineages.) If this autocorrelation is 1, the minimum possible fixation time consistent with the progeny distribution will be realized. If the autocorrelation is 0, selective neutrality governs. Fixation times can be further retarded if the autocorrelation is negative. Note that the autocorrelation of progeny production could be due to cultural factors such as inheritance of wealth or power, as well as genetic factors.
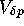 and Coalescence Time
and Coalescence Time
In the case of selective neutrality, the diffusion approximation
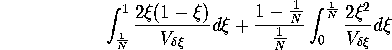
provides that the time until fixation is inversely proportional to the
sampling variance (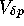 ). Indeed,
). Indeed, 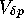 may depend on the
allele frequency p, but if there is a proportionate increase for all
frequencies, the fixation time will be reduced accordingly.
may depend on the
allele frequency p, but if there is a proportionate increase for all
frequencies, the fixation time will be reduced accordingly.
The sampling variance is used to define the (variance) effective population
size (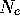 ), and often the effective population size will be specified
rather than the sampling variance. Although both the sampling variance (or
effective population size) and actual population size are needed to evaluate
the above integral, the actual population size is not important if it is
large, hence the formula
), and often the effective population size will be specified
rather than the sampling variance. Although both the sampling variance (or
effective population size) and actual population size are needed to evaluate
the above integral, the actual population size is not important if it is
large, hence the formula 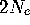 generations for fixation time, which is
really a function of the sampling variance.
generations for fixation time, which is
really a function of the sampling variance.
Autocorrelation and Effective Sampling Variance
If progeny number is independent of genotype, the autocorrelation of progeny
number is the correlation of 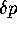 for successive generations.
Fixation does not occur due to the sampling variance in a single generation,
but due to the cumulative effect over many generations. If sampling is
independent between generations, the cumulative variance over n generations
is n times the variance in a single generation, but if the same variation
occurs each generation (complete dependence), the cumulative variance over n
generations is
for successive generations.
Fixation does not occur due to the sampling variance in a single generation,
but due to the cumulative effect over many generations. If sampling is
independent between generations, the cumulative variance over n generations
is n times the variance in a single generation, but if the same variation
occurs each generation (complete dependence), the cumulative variance over n
generations is 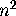 times the variance in a single generation. It can be
shown that if the autocorrelation of number of progeny (or more correctly,
the correlation of
times the variance in a single generation. It can be
shown that if the autocorrelation of number of progeny (or more correctly,
the correlation of 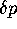 ) separated by n
generations is
) separated by n
generations is 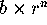 , the cumulative variance over n generations
will be approximately (for large n)
, the cumulative variance over n generations
will be approximately (for large n)
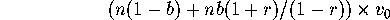
where 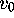 is the variance in a single generation (
is the variance in a single generation (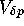 ). Hence
the fixation time (and
). Hence
the fixation time (and 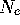 ) will be divided by
) will be divided by 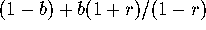 compared to independence between generations.
compared to independence between generations.
A genetic model for a diploid population would have b be the extent to which
progeny number is genetically determined, and 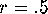 reflecting free
recombination. Cultural or social inheritance models could have b=1 and any
value (less than 1) for r.
reflecting free
recombination. Cultural or social inheritance models could have b=1 and any
value (less than 1) for r.
Effect of Standard Selection
If two alleles with relative viabilities 1+s and 1 are segregating in a
population, the autocorrelation of progeny number between successive
generations (hence correlation of 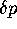 at independent loci) will be
at independent loci) will be
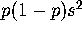 (which is also the genetic variance of the population). This is
a small quantity, hence will not have much effect on coalescent times.
At the locus undergoing selection,
(which is also the genetic variance of the population). This is
a small quantity, hence will not have much effect on coalescent times.
At the locus undergoing selection, 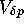 due to selection will be
due to selection will be
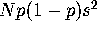 , but in 2N generations (neutral fixation time) the cumulative
effect of the autocorrelation will be less than the effect of drift if
Ns<<1.
, but in 2N generations (neutral fixation time) the cumulative
effect of the autocorrelation will be less than the effect of drift if
Ns<<1.
If variance due to selection is in addition to the background variance, rather
than part of it, it is necessary to assess how much variance selection adds.
Since selection must be manifested as giving entire extra progeny to a few
individuals (rather than fractions of progeny to many), the additional
variance will be 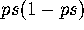 . This will have a greater effect than the
autocorrelation, and if several loci are undergoing selection could be
significant.
. This will have a greater effect than the
autocorrelation, and if several loci are undergoing selection could be
significant.
Summary
The importance of sampling variance has long been acknowledged by its use in calculating the effective population size, but its importance is belied by the impression that it modifies the population size. Rather, the sampling variance is of primary importance for population genetic processes, as demonstrated by the unimportance of the actual population size for fixation times. Differences in sampling variances are very important to the dynamics of population genetics.
Selection as manifested in the autocorrelation of progeny number can be used to define an effective sampling variance. This reflects the increase in the cumulative variance over many generations due to the autocorrelation. The effect of the autocorrelation is definitely subordinate to the underlying sampling variance, but can be significant for population genetic quantities such as coalescent times.
Is Adam Younger than Eve?
A recent estimate of the time since the common ancestral y-chromosome is 270,000 years, while a recent estimate of the time since the common ancestral mitochondrial DNA is at least 436,000 years. Although this discrepancy may be due to the error inherent in such estimates, it is appropriate to ask whether that discrepancy may in fact be real. The y-chromosome is present only in male lineages, and the presence of mitochondria in males does not effect the dynamics in the female population.
The actual size of the male and female populations will be essentially the same due to the nature of the sex determination process. But the population genetics of the male and female population could be quite different. The present focus is to study males and females as haploid populations, rather than to obtain an effective size for a diploid population which incorporates their differences.
Dominant Males and Intense Selection
In many species including humans there is a greater tendency for males to have multiple mates than females. If half the males were withdrawn from the mating pool with multiple mating practiced by the rest, the effective population size, hence coalescent time, of the males would be half that of the females. The observed discrepancy could be real if humans have traditionally had mating practices which are asymmetric with respect to the sexes.
If the autocorrelation along male lineages were 1/3, while there was no
autocorrelation along female lineages, the male coalescent time would be half
that of females. Although 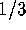 may seem a large value, a survey of 110
students produced a correlation of .45 between the number of siblings the
students had and the number of siblings their fathers had, but only a
correlation of .08 between the number of siblings the students had and the
number of siblings their mothers had. This evidence is only anecdotal, but it
does suggest that selection defined as autocorrelation of progeny number could
explain a twofold difference in coalescent times between the sexes.
may seem a large value, a survey of 110
students produced a correlation of .45 between the number of siblings the
students had and the number of siblings their fathers had, but only a
correlation of .08 between the number of siblings the students had and the
number of siblings their mothers had. This evidence is only anecdotal, but it
does suggest that selection defined as autocorrelation of progeny number could
explain a twofold difference in coalescent times between the sexes.